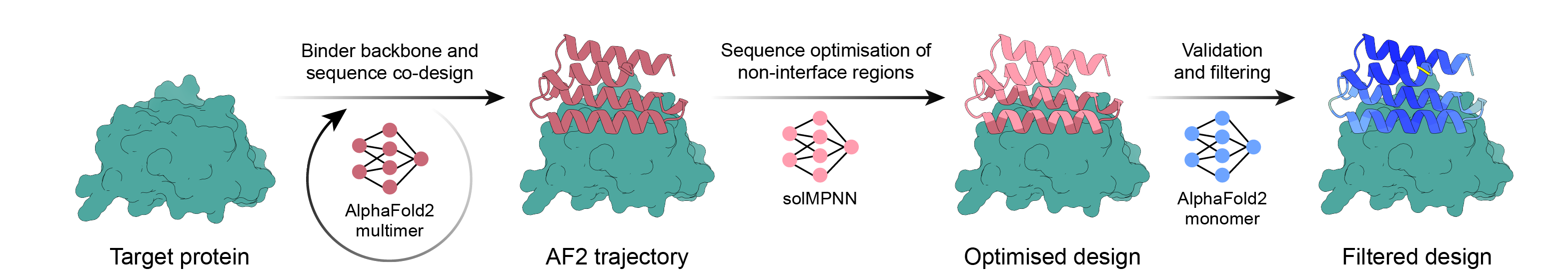
Einfache Binder-Design-Pipeline mit AlphaFold2-Backpropagation, MPNN und PyRosetta. Wählen Sie Ihr Ziel aus und lassen Sie das Skript den Rest der Arbeit erledigen und beenden Sie es, sobald Sie genügend Designs zum Bestellen haben!
Preprint-Link für BindCraft
Zuerst müssen Sie dieses Repository klonen. Ersetzen Sie [install_folder] durch den Pfad, in dem Sie es installieren möchten.
git clone https://github.com/martinpacesa/BindCraft [install_folder]
Navigieren Sie dann mit der CD in Ihren Installationsordner und führen Sie den Installationscode aus. Für die Ausführung von BindCraft ist eine CUDA-kompatible Nvidia-Grafikkarte erforderlich. Bitte geben Sie in der cuda- Einstellung die mit Ihrer Grafikkarte kompatible CUDA-Version an, zum Beispiel „11.8“. Wenn Sie sich nicht sicher sind, lassen Sie das Feld leer, aber es ist möglich, dass bei der Installation die falsche Version ausgewählt wird, was zu Fehlern führt. Geben Sie im pkg_manager an, ob Sie „mamba“ oder „conda“ verwenden. Wenn Sie das Feld leer lassen, wird standardmäßig „conda“ verwendet.
Hinweis: Dieses Installationsskript installiert PyRosetta, für das für kommerzielle Zwecke eine Lizenz erforderlich ist.
bash install_bindcraft.sh --cuda '12.4' --pkg_manager 'conda'
Versuchen Sie immer, die Eingabeziel-PDB auf die kleinstmögliche Größe zu reduzieren! Dadurch wird die Ordnergenerierung erheblich beschleunigt und der GPU-Speicherbedarf minimiert.
Seien Sie bereit, mindestens ein paar Hundert Flugbahnen zu absolvieren, um einige akzeptierte Zielfernrohre zu sehen, bei schwierigen Zielen könnten es sogar ein paar Tausend sein.
Um das Skript lokal auszuführen, müssen Sie zunächst Ihre Ziel-JSON-Datei im Ordner „settings_target“ konfigurieren. In der JSON-Datei sind folgende Einstellungen:
design_path -> path where to save designs and statistics
binder_name -> what to prefix your designed binder files with
starting_pdb -> the path to the PDB of your target protein
chains -> which chains to target in your protein, rest will be ignored
target_hotspot_residues -> which position to target for binder design, for example `1,2-10` or chain specific `A1-10,B1-20` or entire chains `A`, set to null if you want AF2 to select binding site; better to select multiple target residues or a small patch to reduce search space for binder
lengths -> range of binder lengths to design
number_of_final_designs -> how many designs that pass all filters to aim for, script will stop if this many are reached
Führen Sie dann das Binder-Design-Skript aus:
sbatch ./bindcraft.slurm --settings './settings_target/PDL1.json' --filters './settings_filters/default_filters.json' --advanced './settings_advanced/default_4stage_multimer.json'
Das Einstellungsflag sollte auf Ihre Ziel-.json-Datei verweisen, die Sie oben festgelegt haben. Das Filter- Flag zeigt auf den JSON, in dem die Designfilter angegeben sind (Standard ist ./filters/default_filters.json). Das Flag „Erweitert“ verweist auf Ihre erweiterten Einstellungen (Standard ist ./advanced_settings/default_4stage_multimer.json). Wenn Sie die Filter und erweiterten Einstellungsflags weglassen, wird automatisch auf die Standardeinstellungen verwiesen.
Wenn Ihr Computer SLURM nicht unterstützt, können Sie alternativ den Code direkt ausführen, indem Sie die Umgebung in Conda aktivieren und den Python-Code ausführen:
conda activate BindCraft
cd /path/to/bindcraft/folder/
python -u ./bindcraft.py --settings './settings_target/PDL1.json' --filters './settings_filters/default_filters.json' --advanced './settings_advanced/default_4stage_multimer.json'
Wir empfehlen, mindestens 100 endgültige Designs zu erstellen, die alle Filter bestehen, und dann die besten 5–20 für die experimentelle Charakterisierung zu bestellen. Wenn Bindungen mit hoher Affinität erforderlich sind, ist es besser, mehr zu screenen, da die für die Rangfolge verwendete ipTM-Metrik kein guter Prädiktor für die Affinität ist, sich aber als guter binärer Prädiktor für die Bindung erwiesen hat.
Nachfolgend finden Sie Erläuterungen zu einzelnen Filtern und erweiterten Einstellungen.
Hier sind die erweiterten Einstellungen, die den Designprozess steuern:
omit_AAs -> which amino acids to exclude from design (note: they can still occur if no other options are possible in the position)
force_reject_AA -> whether to force reject design if it contains any amino acids specified in omit_AAs
design_algorithm -> which design algorithm for the trajecory to use, the currently implemented algorithms are below
use_multimer_design -> whether to use AF2-ptm or AF2-multimer for binder design; the other model will be used for validation then
num_recycles_design -> how many recycles of AF2 for design
num_recycles_validation -> how many recycles of AF2 use for structure prediction and validation
sample_models = True -> whether to randomly sample parameters from AF2 models, recommended to avoid overfitting
rm_template_seq_design -> remove target template sequence for design (increases target flexibility)
rm_template_seq_predict -> remove target template sequence for reprediction (increases target flexibility)
rm_template_sc_design -> remove sidechains from target template for design
rm_template_sc_predict -> remove sidechains from target template for reprediction
# Design iterations
soft_iterations -> number of soft iterations (all amino acids considered at all positions)
temporary_iterations -> number of temporary iterations (softmax, most probable amino acids considered at all positions)
hard_iterations -> number of hard iterations (one hot encoding, single amino acids considered at all positions)
greedy_iterations -> number of iterations to sample random mutations from PSSM that reduce loss
greedy_percentage -> What percentage of protein length to mutate during each greedy iteration
# Design weights, higher value puts more weight on optimising the parameter.
weights_plddt -> Design weight - pLDDT of designed chain
weights_pae_intra -> Design weight - PAE within designed chain
weights_pae_inter -> Design weight - PAE between chains
weights_con_intra -> Design weight - maximise number of contacts within designed chain
weights_con_inter -> Design weight - maximise number of contacts between chains
intra_contact_distance -> Cbeta-Cbeta cutoff distance for contacts within the binder
inter_contact_distance -> Cbeta-Cbeta cutoff distance for contacts between binder and target
intra_contact_number -> how many contacts each contact esidue should make within a chain, excluding immediate neighbours
inter_contact_number -> how many contacts each contact residue should make between chains
weights_helicity -> Design weight - helix propensity of the design, Default 0, negative values bias towards beta sheets
random_helicity -> whether to randomly sample helicity weights for trajectories, from -1 to 1
# Additional losses
use_i_ptm_loss -> Use i_ptm loss to optimise for interface pTM score?
weights_iptm -> Design weight - i_ptm between chains
use_rg_loss -> use radius of gyration loss?
weights_rg -> Design weight - radius of gyration weight for binder
use_termini_distance_loss -> Try to minimise distance between N- and C-terminus of binder? Helpful for grafting
weights_termini_loss -> Design weight - N- and C-terminus distance minimisation weight of binder
# MPNN settings
mpnn_fix_interface -> whether to fix the interface designed in the starting trajectory
num_seqs -> number of MPNN generated sequences to sample and predict per binder
max_mpnn_sequences -> how many maximum MPNN sequences per trajectory to save if several pass filters
max_tm-score_filter -> filter out final lower ranking designs by this TM score cut off relative to all passing designs
max_seq-similarity_filter -> filter out final lower ranking designs by this sequence similarity cut off relative to all passing designs
sampling_temp = 0.1 -> sampling temperature for amino acids, T=0.0 means taking argmax, T>>1.0 means sampling randomly.")
# MPNN settings - advanced
sample_seq_parallel -> how many sequences to sample in parallel, reduce if running out of memory
backbone_noise -> backbone noise during sampling, 0.00-0.02 are good values
model_path -> path to the MPNN model weights
mpnn_weights -> whether to use "original" mpnn weights or "soluble" weights
save_mpnn_fasta -> whether to save MPNN sequences as fasta files, normally not needed as the sequence is also in the CSV file
# AF2 design settings - advanced
num_recycles_design -> how many recycles of AF2 for design
num_recycles_validation -> how many recycles of AF2 use for structure prediction and validation
optimise_beta -> optimise predictions if beta sheeted trajectory detected?
optimise_beta_extra_soft -> how many extra soft iterations to add if beta sheets detected
optimise_beta_extra_temp -> how many extra temporary iterations to add if beta sheets detected
optimise_beta_recycles_design -> how many recycles to do during design if beta sheets detected
optimise_beta_recycles_valid -> how many recycles to do during reprediction if beta sheets detected
# Optimise script
remove_unrelaxed_trajectory -> remove the PDB files of unrelaxed designed trajectories, relaxed PDBs are retained
remove_unrelaxed_complex -> remove the PDB files of unrelaxed predicted MPNN-optimised complexes, relaxed PDBs are retained
remove_binder_monomer -> remove the PDB files of predicted binder monomers after scoring to save space
zip_animations -> at the end, zip Animations trajectory folder to save space
zip_plots -> at the end, zip Plots trajectory folder to save space
save_trajectory_pickle -> save pickle file of the generated trajectory, careful, takes up a lot of storage space!
max_trajectories -> how many maximum trajectories to generate, for benchmarking
acceptance_rate -> what fraction of trajectories should yield designs passing the filters, if the proportion of successful designs is less than this fraction then the script will stop and you should adjust your design weights
start_monitoring -> after what number of trajectories should we start monitoring acceptance_rate, do not set too low, could terminate prematurely
# debug settings
enable_mpnn = True -> whether to enable MPNN design
enable_rejection_check -> enable rejection rate check
Hier sind die Funktionen, nach denen Ihre Designs gefiltert werden. Wenn Sie einige nicht verwenden möchten, legen Sie einfach Null als Schwellenwert fest. Die Option „höher“ gibt an, ob Werte über dem Schwellenwert beibehalten (wahr) oder niedriger (falsch) werden sollen. Merkmale, die mit N_ beginnen, entsprechen den Statistiken für jedes AlphaFold-Modell. Die Durchschnittswerte gelten für alle vorhergesagten Modelle.
MPNN_score -> MPNN sequence score, generally not recommended as it depends on protein
MPNN_seq_recovery -> MPNN sequence recovery of original trajectory
pLDDT -> pLDDT confidence score of AF2 complex prediction, normalised to 0-1
pTM -> pTM confidence score of AF2 complex prediction, normalised to 0-1
i_pTM -> interface pTM confidence score of AF2 complex prediction, normalised to 0-1
pAE -> predicted alignment error of AF2 complex prediction, normalised compared AF2 by n/31 to 0-1
i_pAE -> predicted interface alignment error of AF2 complex prediction, normalised compared AF2 by n/31 to 0-1
i_pLDDT -> interface pLDDT confidence score of AF2 complex prediction, normalised to 0-1
ss_pLDDT -> secondary structure pLDDT confidence score of AF2 complex prediction, normalised to 0-1
Unrelaxed_Clashes -> number of interface clashes before relaxation
Relaxed_Clashes -> number of interface clashes after relaxation
Binder_Energy_Score -> Rosetta energy score for binder alone
Surface_Hydrophobicity -> surface hydrophobicity fraction for binder
ShapeComplementarity -> interface shape complementarity
PackStat -> interface packstat rosetta score
dG -> interface rosetta dG energy
dSASA -> interface delta SASA (size)
dG/dSASA -> interface energy divided by interface size
Interface_SASA_% -> Fraction of binder surface covered by the interface
Interface_Hydrophobicity -> Interface hydrophobicity fraction of binder interface
n_InterfaceResidues -> number of interface residues
n_InterfaceHbonds -> number of hydrogen bonds at the interface
InterfaceHbondsPercentage -> number of hydrogen bonds compared to interface size
n_InterfaceUnsatHbonds -> number of unsatisfied buried hydrogen bonds at the interface
InterfaceUnsatHbondsPercentage -> number of unsatisfied buried hydrogen bonds compared to interface size
Interface_Helix% -> proportion of alfa helices at the interface
Interface_BetaSheet% -> proportion of beta sheets at the interface
Interface_Loop% -> proportion of loops at the interface
Binder_Helix% -> proportion of alfa helices in the binder structure
Binder_BetaSheet% -> proportion of beta sheets in the binder structure
Binder_Loop% -> proportion of loops in the binder structure
InterfaceAAs -> number of amino acids of each type at the interface
HotspotRMSD -> unaligned RMSD of binder compared to original trajectory, in other words how far is binder in the repredicted complex from the original binding site
Target_RMSD -> RMSD of target predicted in context of the designed binder compared to input PDB
Binder_pLDDT -> pLDDT confidence score of binder predicted alone
Binder_pTM -> pTM confidence score of binder predicted alone
Binder_pAE -> predicted alignment error of binder predicted alone
Binder_RMSD -> RMSD of binder predicted alone compared to original trajectory
Vielen Dank an Lennart Nickel, Yehlin Cho, Casper Goverde und Sergey Ovchinnikov für ihre Hilfe beim Codieren und Besprechen von Ideen. Dieses Repository verwendet Code von: